DireX - Low-resolution Structure Refinement |
DireX performs efficient geometry-based conformational sampling of protein structures under experimental restraints. It combines prior structural information with experimental data through the Deformable Elastic Network (DEN) approach which drastically reduces over-fitting.
Experimental restraints which can be used in the current version are:
- (Electron) Density Map (obtained e.g. from X-ray crystallography, Electron microscopy, or SAXS)
- Distance Restraints (e.g. from NMR or FRET experiments)
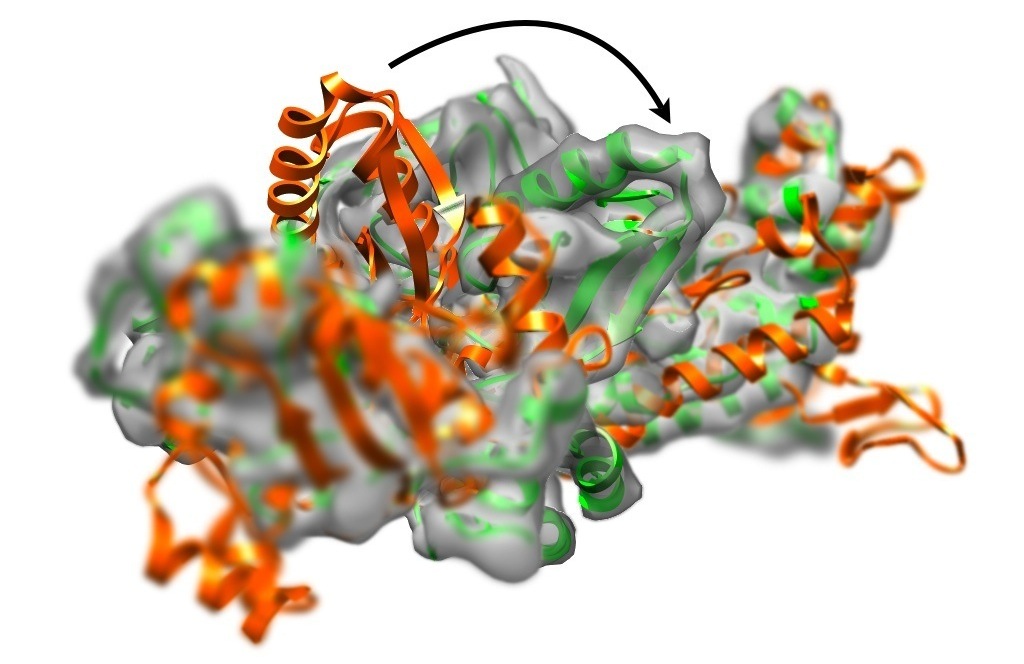
To get a first impression: Watch a movie of fitting Elongation factor 2 (EF-2) into an 8 Å density map.
Aug 8, 2019: New DireX version released (0.7.1):
Added a new turbo mode (use command line argument -turbo), which prevents computing a new model map at each step. The mode uses only the target map as a restraint as does not optimize the overlap of model map and target map. This turbo mode target function is more similar to what is used in MDFF.